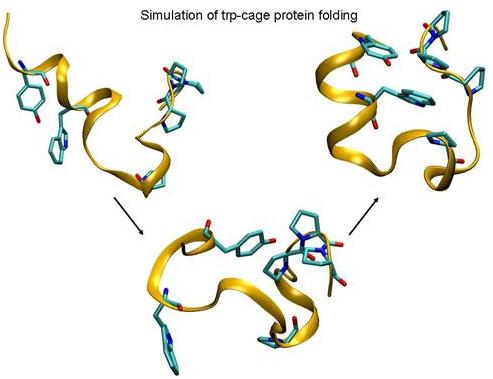
IJMS | Free Full-Text | Folding of Trp-cage Mini Protein Using Temperature and Biasing Potential Replica—Exchange Molecular Dynamics Simulations

RCSB PDB - 2M7C: Circular Permutation of the Trp-cage: Fold Rescue upon Addition of a Hydrophobic Staple

Crystal and NMR structures of a Trp-cage mini-protein benchmark for computational fold prediction | PNAS
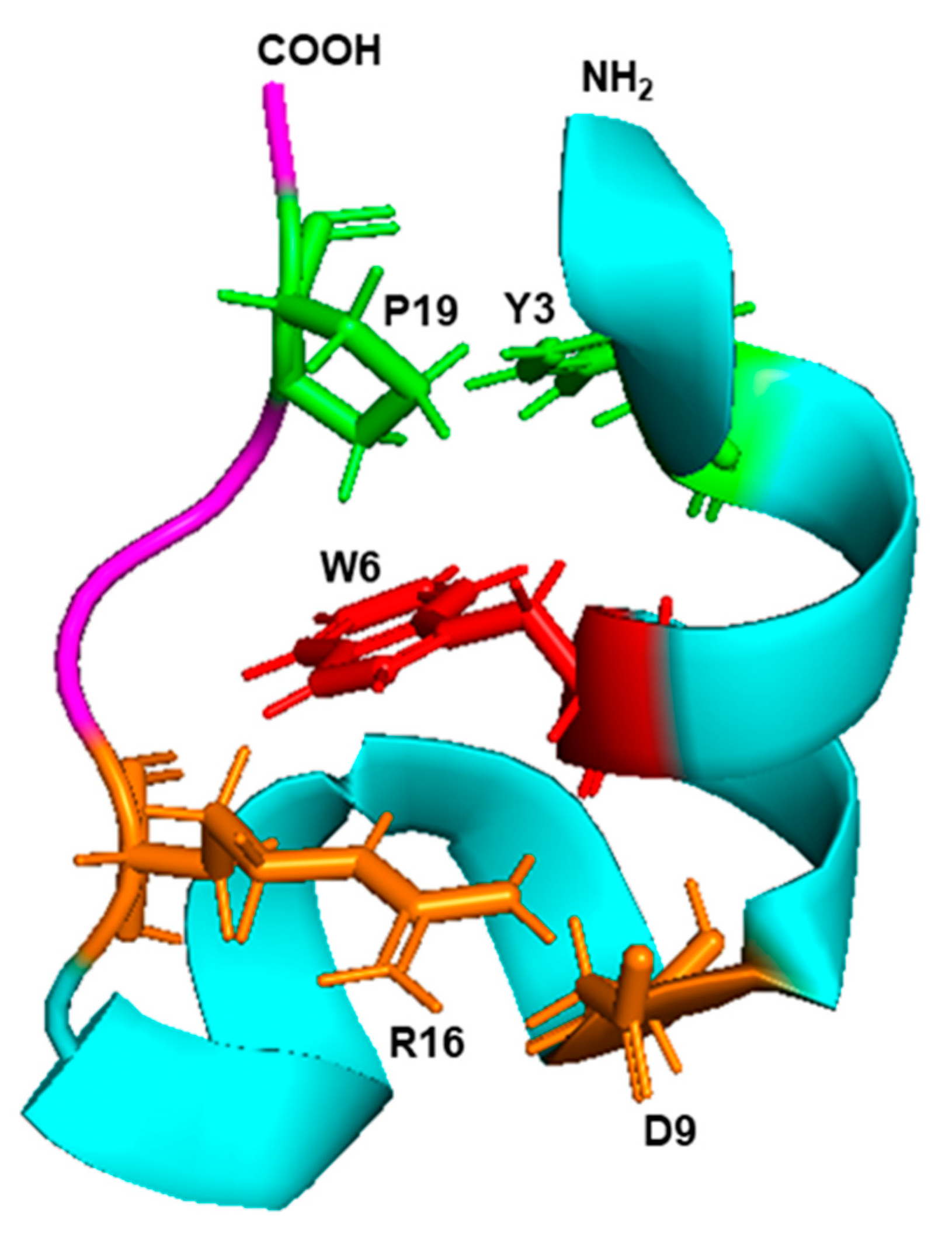
IJMS | Free Full-Text | The Uniqueness of Tryptophan in Biology: Properties, Metabolism, Interactions and Localization in Proteins

Folding/unfolding of protein Trp cage in aqueous osmolyte solutions under polar confinement - ScienceDirect

Cooperation between a Salt Bridge and the Hydrophobic Core Triggers Fold Stabilization in a Trp-Cage Miniprotein | Biochemistry
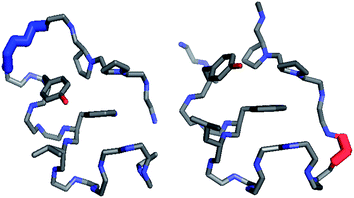
Circular permutation of the Trp-cage: fold rescue upon addition of a hydrophobic staple - RSC Advances (RSC Publishing) DOI:10.1039/C3RA43674H

Folding Trp-Cage to NMR Resolution Native Structure Using a Coarse-Grained Protein Model: Biophysical Journal

The Trp Cage Motif as a Scaffold for the Display of a Randomized Peptide Library on Bacteriophage T7 - ScienceDirect
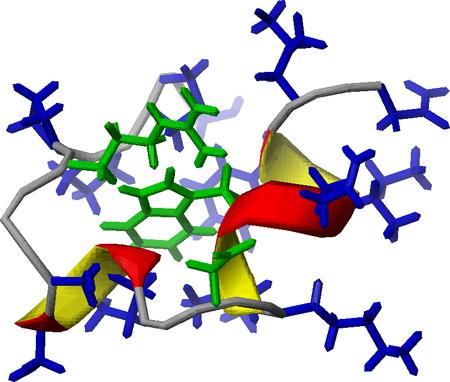
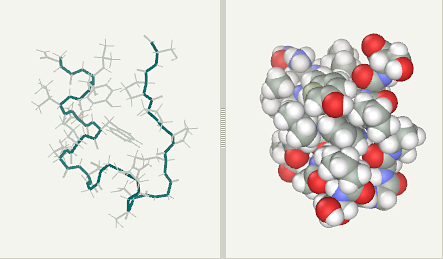
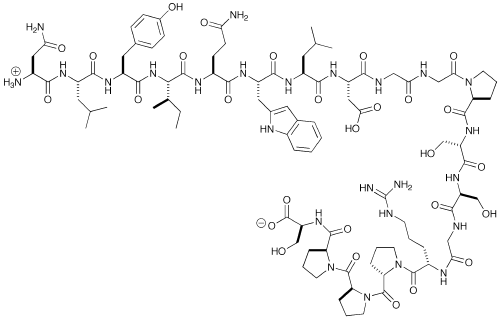
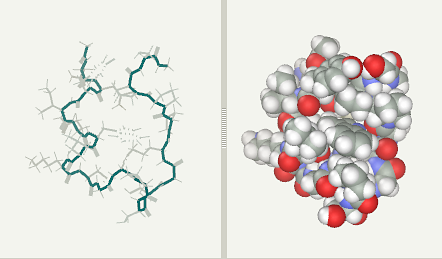
Kutatás - Kutatásaink - NMR-spectroscopy - Design and characterization of Trp-cage miniproteins - MTA-ELTE Fehérjemodellező Kutatócsoport és Szerkezeti Kémia és Biológia Laboratórium
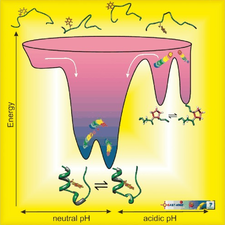
Kutatás - Kutatásaink - NMR-spectroscopy - Design and characterization of Trp-cage miniproteins - MTA-ELTE Fehérjemodellező Kutatócsoport és Szerkezeti Kémia és Biológia Laboratórium

.jpg)