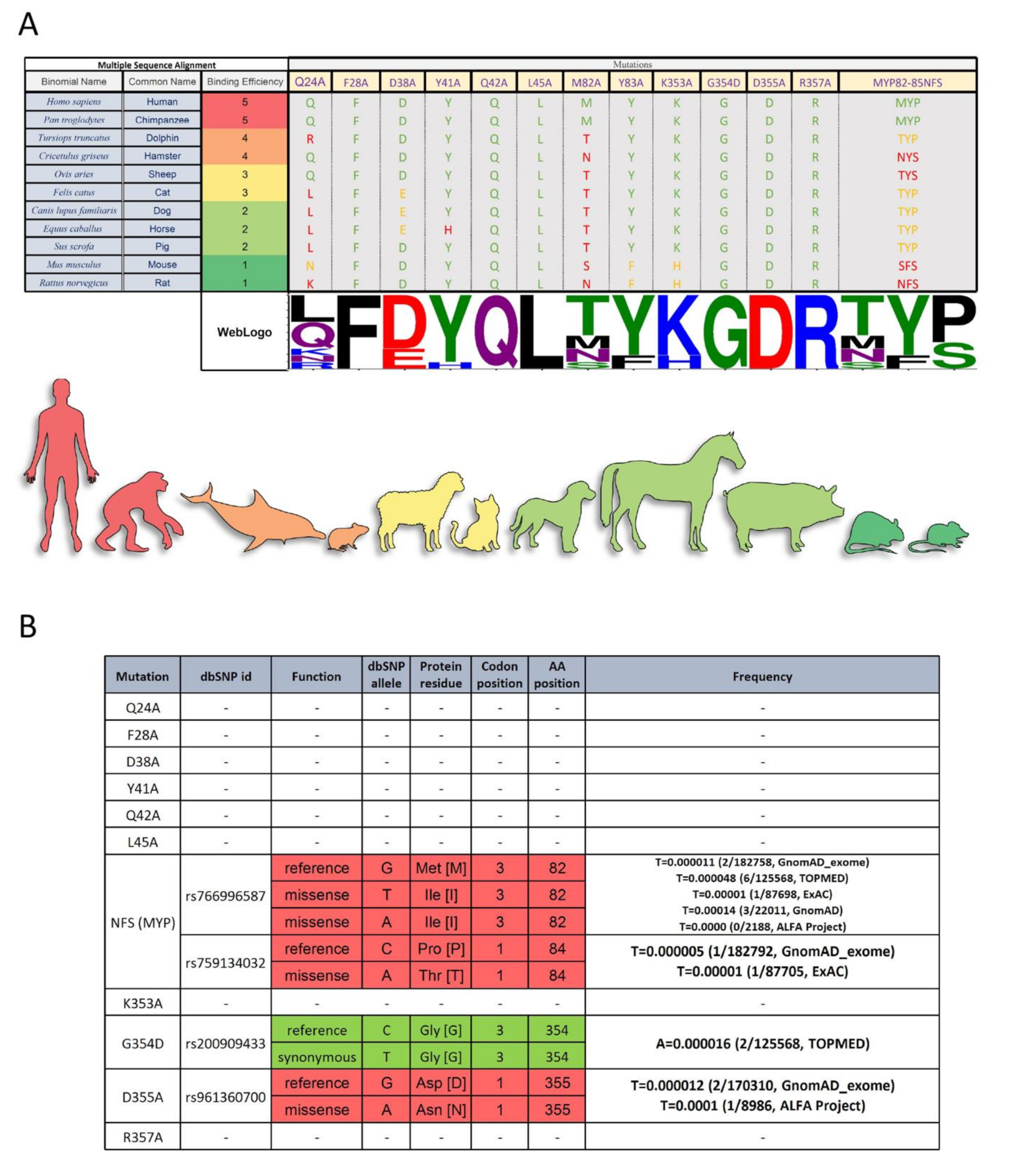
IJMS | Free Full-Text | Characterization of Critical Determinants of ACE2–SARS CoV-2 RBD Interaction | HTML
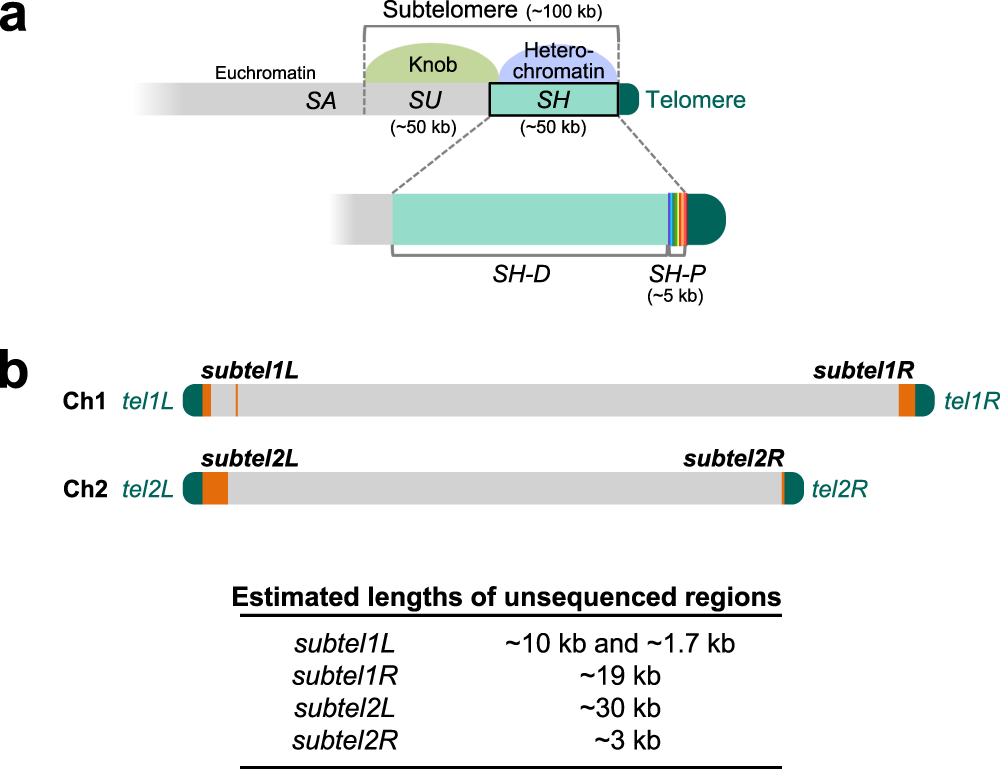
Complete sequences of Schizosaccharomyces pombe subtelomeres reveal multiple patterns of genome variation | Nature Communications

Identification of errors introduced during high throughput sequencing of the T cell receptor repertoire | BMC Genomics | Full Text

Sequence and structural characteristics of circRNA-1926 in cashmere... | Download Scientific Diagram

Enrichment of low-frequency functional variants revealed by whole-genome sequencing of multiple isolated European populations | Nature Communications

MUSCLE: a multiple sequence alignment method with reduced time and space complexity | BMC Bioinformatics | Full Text

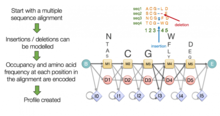
The flowchart of generating the profile-based protein sequences. The... | Download Scientific Diagram
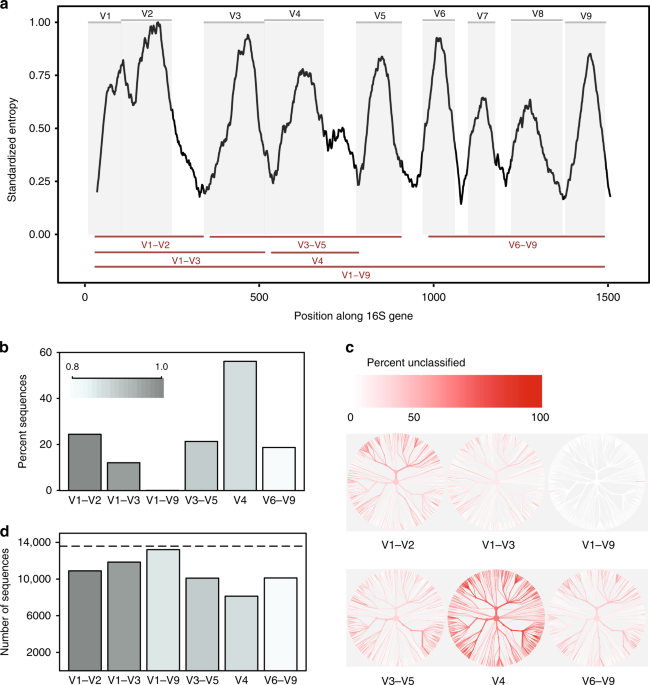
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications
PLOS ONE: Variable Frequency of Plastid RNA Editing among Ferns and Repeated Loss of Uridine-to-Cytidine Editing from Vascular Plants
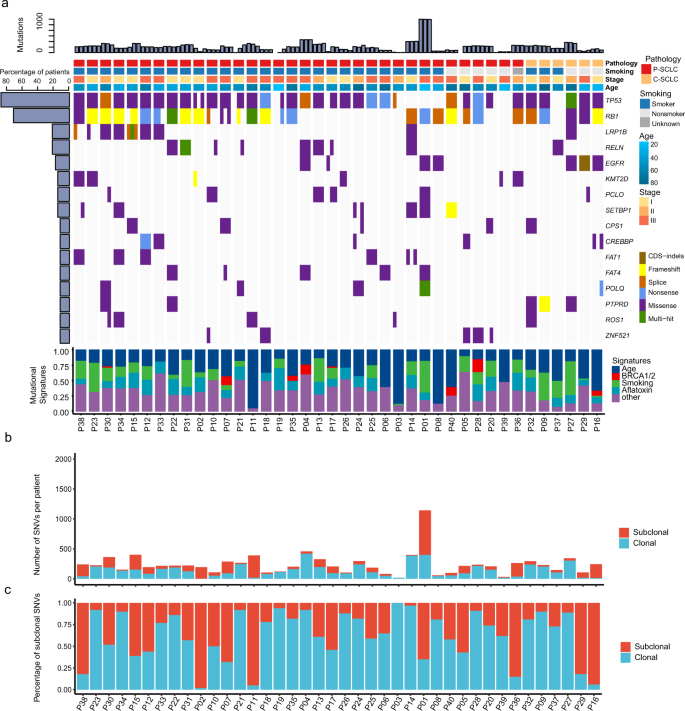
Multi-region exome sequencing reveals the intratumoral heterogeneity of surgically resected small cell lung cancer | Nature Communications

Application of multiple sequence alignment profiles to improve protein secondary structure prediction - Cuff - 2000 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library

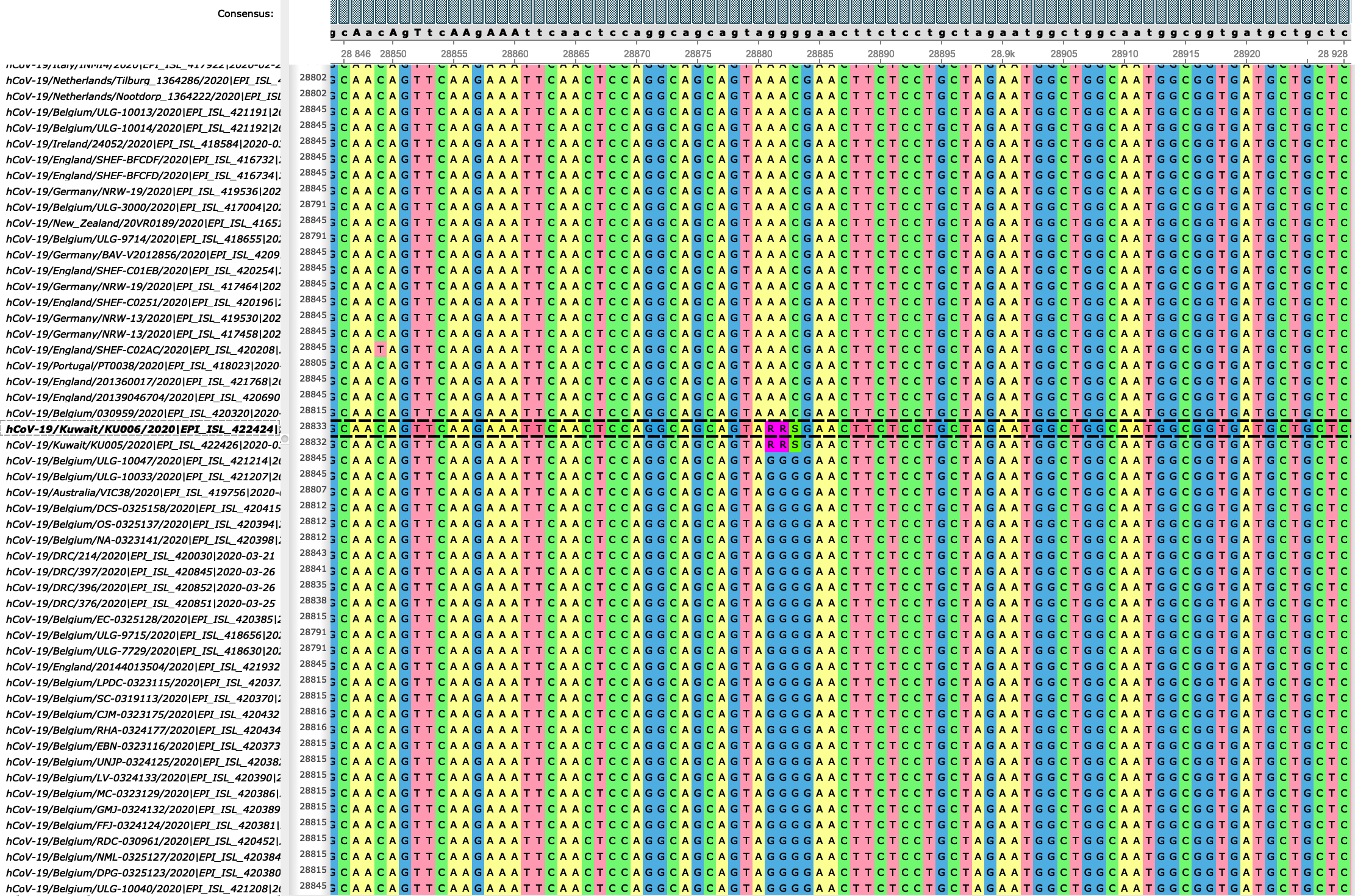
Distribution of SARS-CoV-2 SNPs based on multiple sequence alignment of... | Download Scientific Diagram