
The GAP arginine finger movement into the catalytic site of Ras increases the activation entropy | PNAS

Ras and GTPase-activating protein (GAP) drive GTP into a precatalytic state as revealed by combining FTIR and biomolecular simulations | PNAS

GTP Hydrolysis Is Not Important for Ypt1 GTPase Function in Vesicular Transport | Molecular and Cellular Biology

See previous page) (A) rates of intrinsic and GAP-mediated hydrolysis... | Download Scientific Diagram

Plexins Are GTPase-Activating Proteins for Rap and Are Activated by Induced Dimerization | Science Signaling

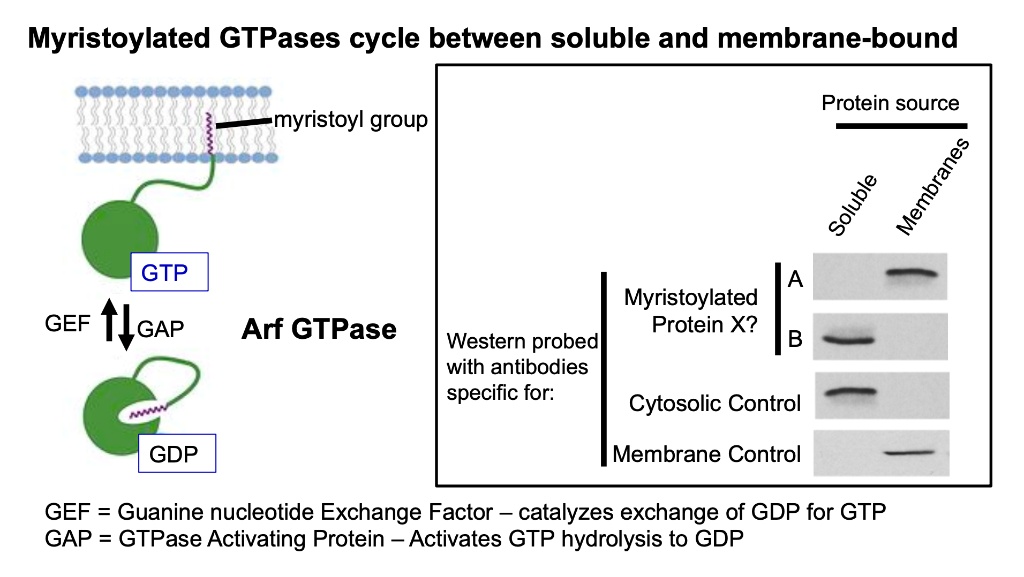
Regulation of GTP Hydrolysis on ADP-ribosylation Factor-1 at the Golgi Membrane* - Journal of Biological Chemistry
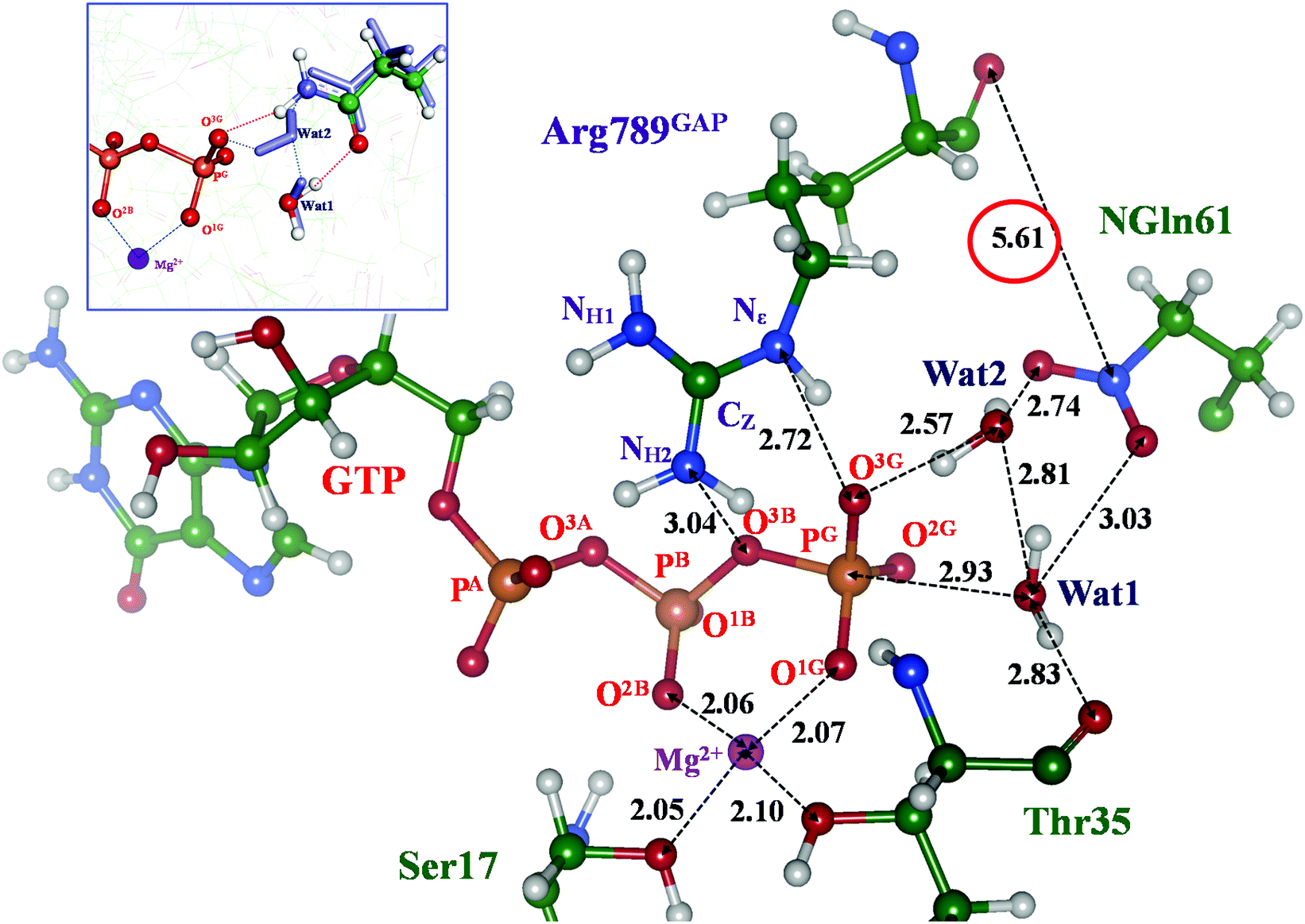
Diversity of mechanisms in Ras–GAP catalysis of guanosine triphosphate hydrolysis revealed by molecular modeling - Organic & Biomolecular Chemistry (RSC Publishing) DOI:10.1039/C9OB00463G

Structural plasticity mediates distinct GAP‐dependent GTP hydrolysis mechanisms in Rab33 and Rab5 - Majumdar - 2017 - The FEBS Journal - Wiley Online Library

Molecules | Free Full-Text | Structural Insights into the Regulation Mechanism of Small GTPases by GEFs | HTML

The GAP arginine finger movement into the catalytic site of Ras increases the activation entropy | PNAS

Structural and Functional Analysis of the ARF1–ARFGAP Complex Reveals a Role for Coatomer in GTP Hydrolysis: Cell

The energetics of the GTP hydrolysis in the Gln61Leu mutant of Ras/... | Download Scientific Diagram










![PDF] Modeling the mechanisms of biological GTP hydrolysis. | Semantic Scholar PDF] Modeling the mechanisms of biological GTP hydrolysis. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d73a3c179ccd59a0d10afd8fcec901434302c963/2-Figure1-1.png)

