Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv
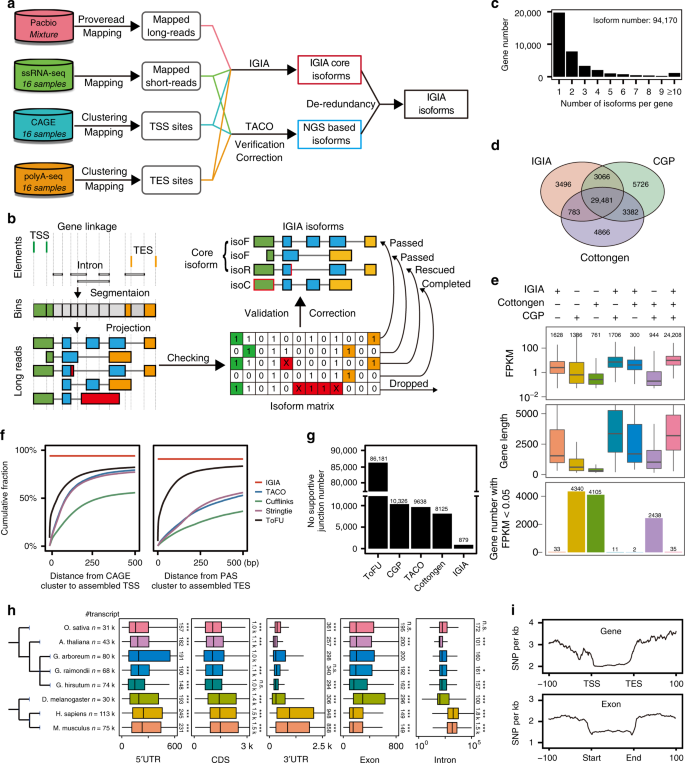
Multi-strategic RNA-seq analysis reveals a high-resolution transcriptional landscape in cotton | Nature Communications
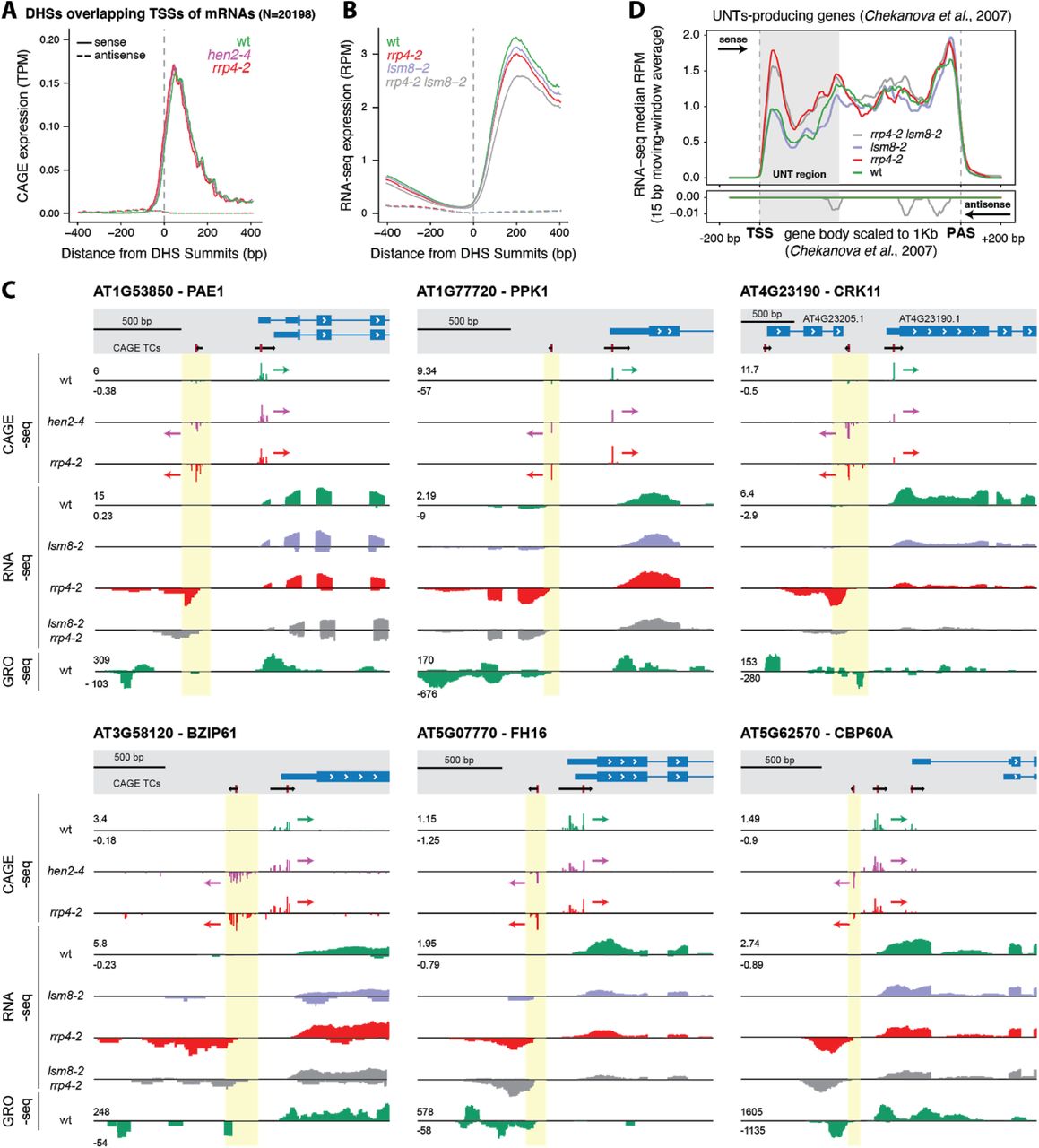
Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv

Cap analysis of gene expression reveals alternative promoter usage in a rat model of hypertension | Life Science Alliance

Nuclear stability and transcriptional directionality separate functionally distinct RNA species | Nature Communications

Maintenance of Normal Blood Pressure and Renal Functions Are Independent Effects of Angiotensin-converting Enzyme* - Journal of Biological Chemistry
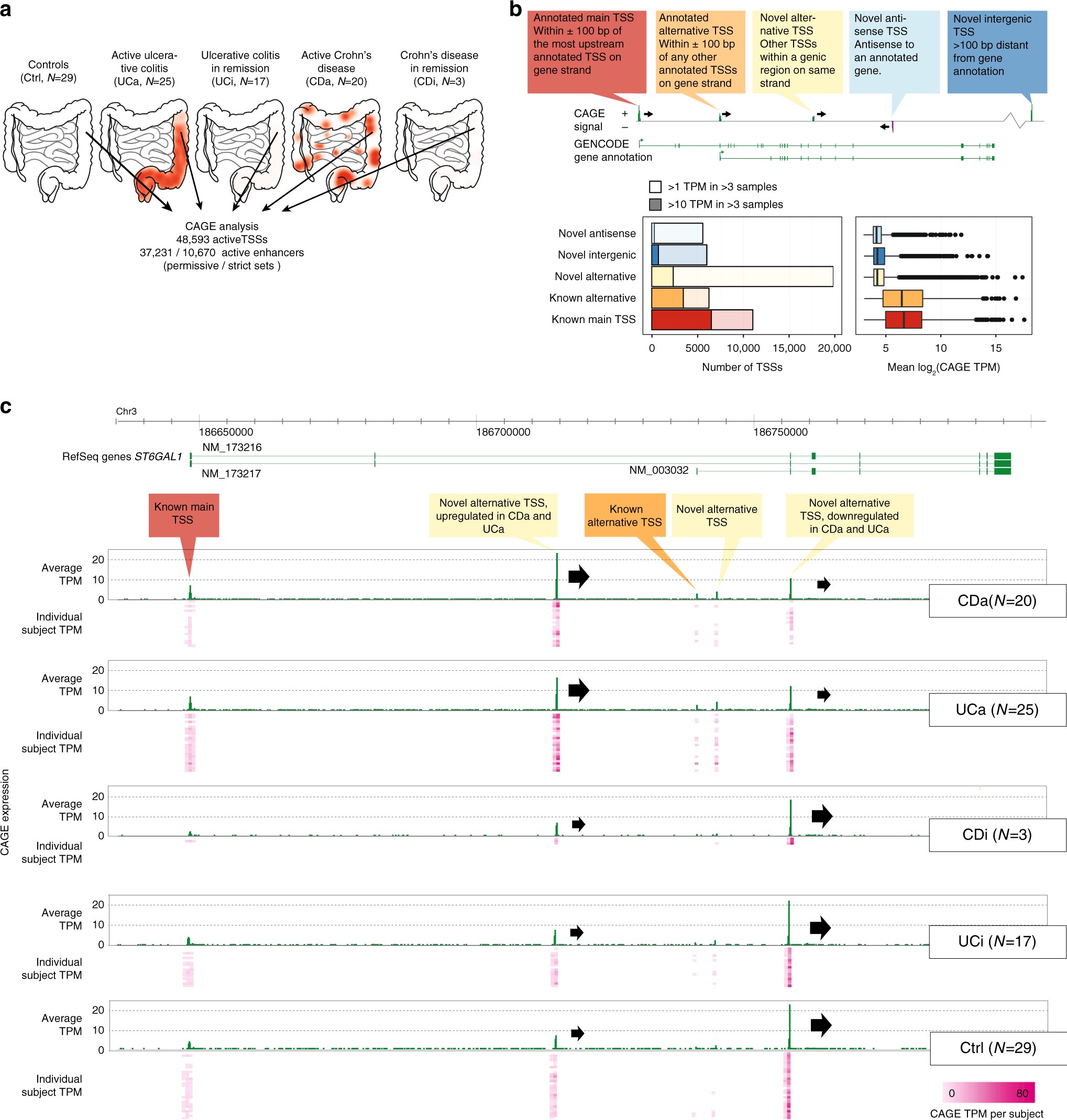
Characterization of the enhancer and promoter landscape of inflammatory bowel disease from human colon biopsies | Nature Communications
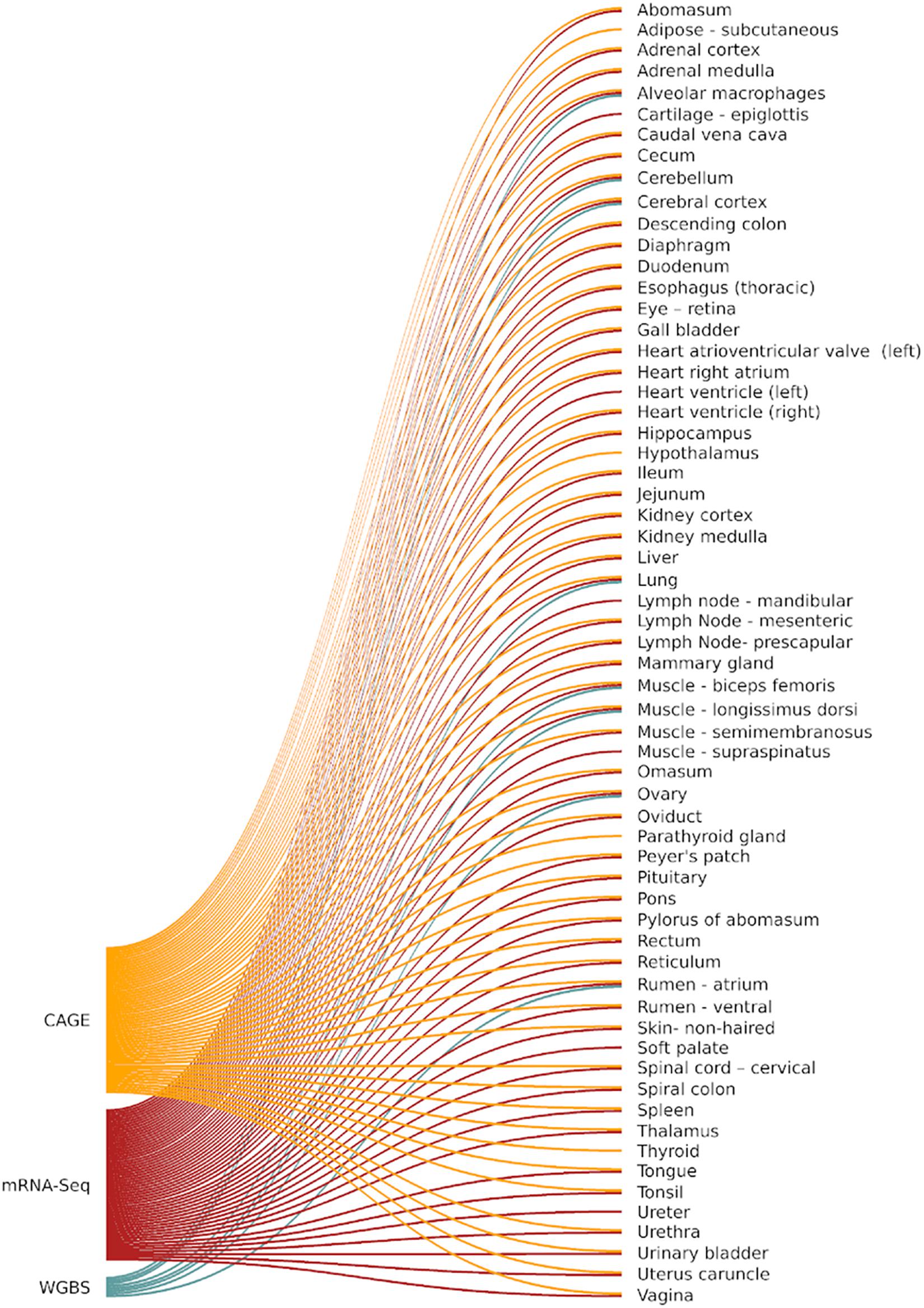
Frontiers | Global Analysis of Transcription Start Sites in the New Ovine Reference Genome (Oar rambouillet v1.0) | Genetics
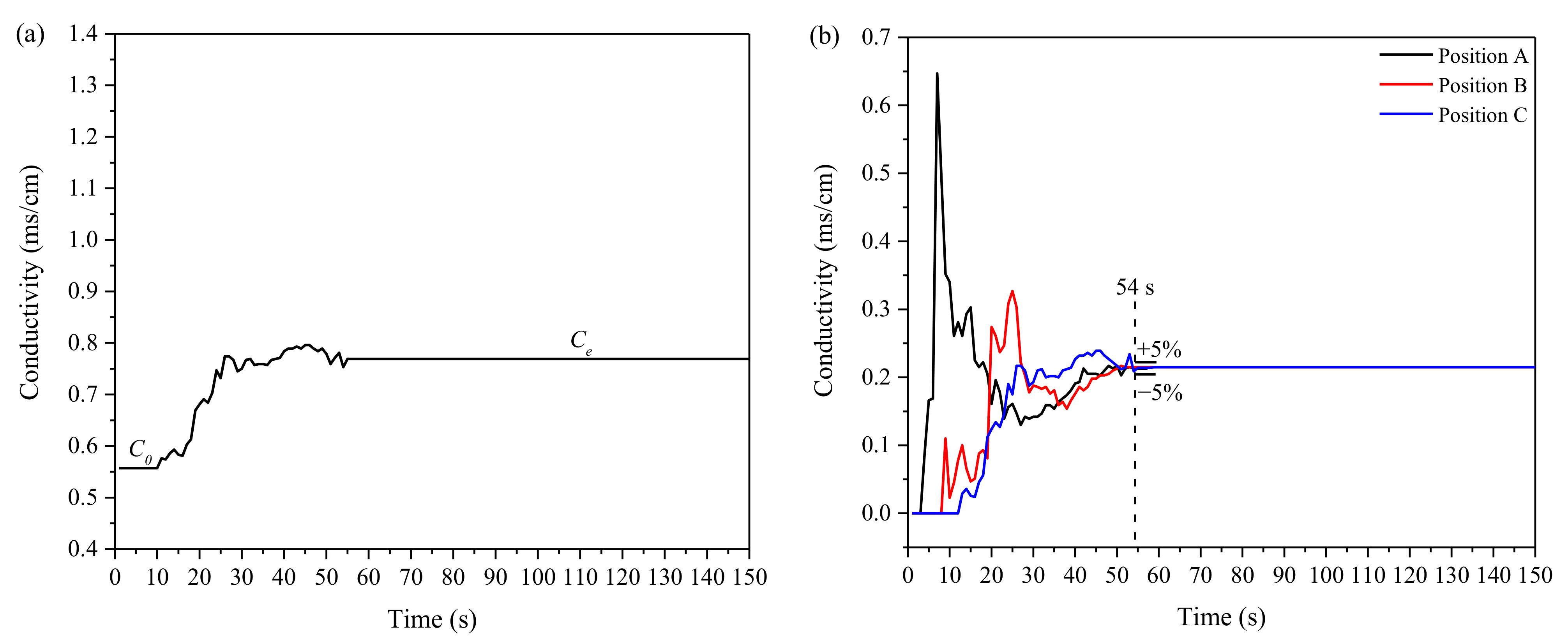
Metals | Free Full-Text | Effect of Side Blowing on Fluid Flow and Mixing Phenomenon in Gas-Stirred Ladle | HTML

TATA and paused promoters active in differentiated tissues have distinct expression characteristics | Molecular Systems Biology

TATA and paused promoters active in differentiated tissues have distinct expression characteristics | Molecular Systems Biology
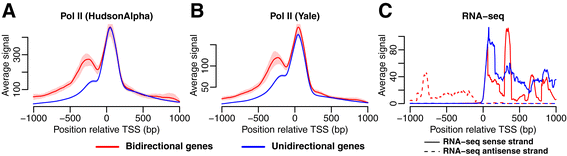
Different distribution of histone modifications in genes with unidirectional and bidirectional transcription and a role of CTCF and cohesin in directing transcription | BMC Genomics | Full Text

Multiple transcription initiation and alternative promoter usage in G.... | Download Scientific Diagram

Features of eSTARR-seq enhancers a, Scatterplot of activity vs GRO-cap... | Download Scientific Diagram

Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv
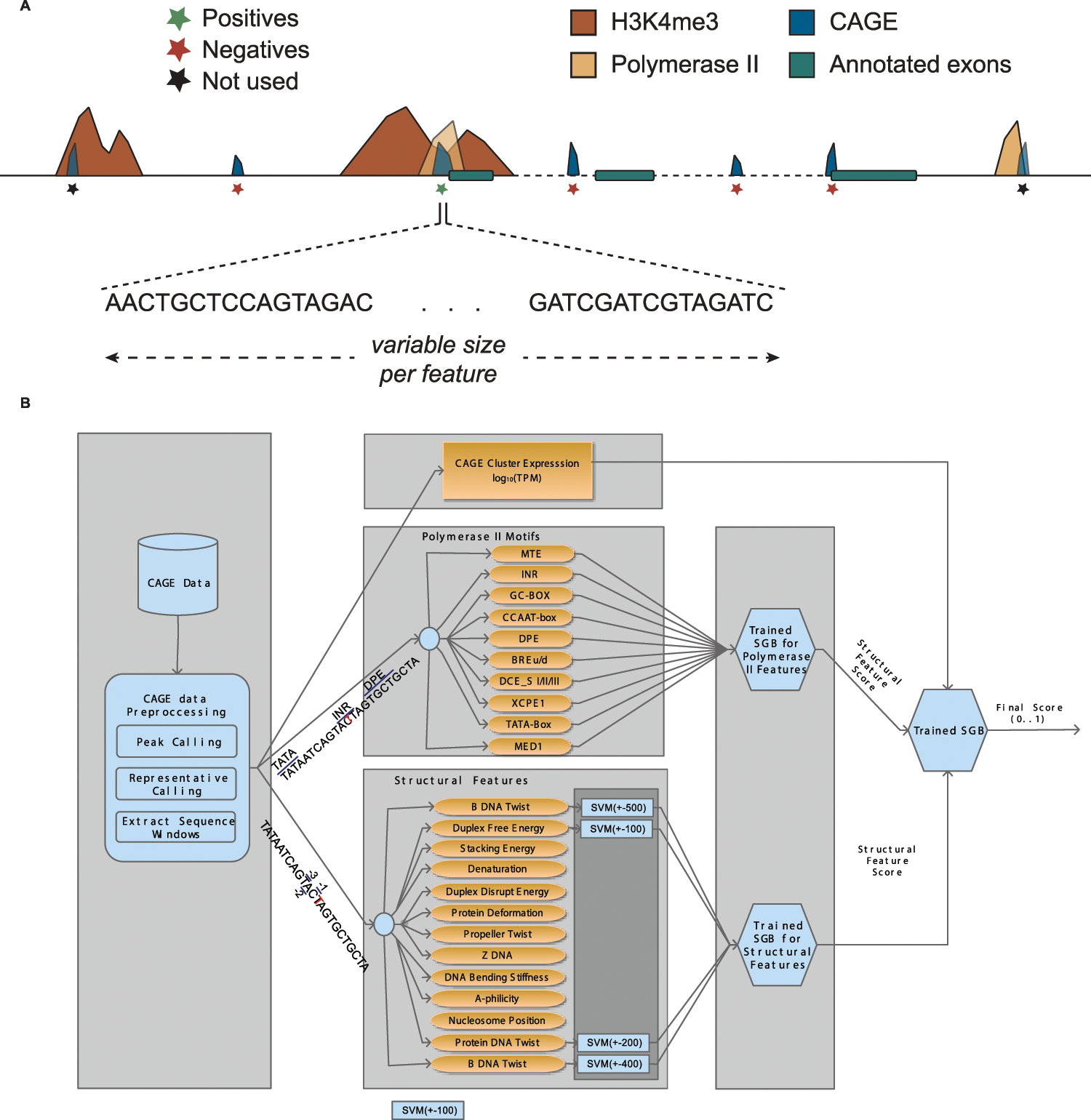
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports